|
 |


- Search
| Int Neurourol J > Volume 26(Suppl 2); 2022 > Article |
|
ABSTRACT
Purpose
In Alzheimer disease (AD), brain regions such as the cortex and the hippocampus show abundant amyloid load which correlates with cognitive function decline. Prior to the significant development of AD pathophysiology, patients report the manifestation of neuropsychiatric symptoms, indicating a functional interplay between basal ganglia structures and hippocampal regions. Zinc finger and BTB domain-containing protein 16 (ZBTB16) is a transcription factor that controls the expression of downstream genes and the involvement of ZBTB16 in the striatum undergoing pathological aging in AD and the resulting behavioral phenotypes has not yet been explored.
Methods
To study molecular alterations in AD pathogenesis, we analyzed the brain from amyloid precursor protein (APP)/ presenilin 1 (PS1) transgenic mice. The molecular changes in the striatal region of the brain were analyzed via the immunoblotting, and the quantitative RNA sequencing. The cognitive impairments of APP/PS1 mice were assessed via 3 behavioral tests: 3-chamber test, Y-maze test, and noble object recognition test. And multielectrode array experiments for the analysis of the neuronal activity of the striatum in APP/PS1 mice was performed.
Results
We found that the alteration in ZBTB16 levels that occurred in the early ages of the pathologically aging striatum coalesces with the disruption of transcriptional dysregulation while causing social memory deficits, anxiety-like behavior. The early ZBTB16 knockdown treatment in the striatum of APP/PS1 mice rescued cognition that continued into later age.
- Based on the results from the striatal transcriptome analysis, the functional role of the ZBTB16 was studied.
- As a result of the in-silico analysis, the ZBTB16 could be analyzed one of the binding target of the MeCP2.
- The knock-down of the increased ZBTB16 led to the rescue phenotypes in the cognitive behaviors and the neuronal activity of the striatum.
Dysregulated gene expression in the brain may lead to neuronal dysfunction in neurodegenerative diseases. In the case of Alzheimer disease (AD), numerous pathogenic factors causing or exacerbating the disease could induce changes in the gene expression profiles of the brain [1]. Through recent studies, functional regulators of gene expression related to the pathogenesis of the AD have been reported [2]. Identifying the pivotal regulator proteins of gene expression such as specific transcription factors leading to transcriptomic changes will characterize more comprehensive target of the AD treatment.
In particular, Zinc finger and BTB domain-containing protein 16 (ZBTB16) has been found to play multiple roles as a transcription factor and an autophagy regulator, respectively. As a transcription factor, the role of ZBTB16 in determining dopaminergic cell fate has been reported with its zinc finger domain that is involved in controlling the expression of the downstream genes [3]. At the same time, as an autophagy regulator, the E3 ligase complex activation by ZBTB16 leads to the degradation of ATG14L which in turn may inhibit autophagy [4]. Although previous findings show the mechanistic role of striatal ZBTB16 in the scope of AD and Huntington disease, exact phenotypic characteristics of the direct manipulation of ZBTB16 in the striatum of AD mice have yet to be elucidated. In previous study, we elucidated the epigenetic dysregulation of the striatal MeCP2 in AD pathogenesis [5]. Prior to our report, a number of papers revealed that changes in the DNA methylation and the epigenetic dysregulation of MeCP2 were related with changes in the expression level of ZBTB16 [6,7]. Additionally, alterations in DNA methylation by the ZBTB16 could also regulate the DNA occupancy of MeCP2 [8]. Based on this, we hypothesized that the function of ZBTB16 as a downstream target of the epigenetic dysregulation of MeCP2 might be involved in AD pathogenesis. In this study, we aimed to uncover the behavioral changes occurred by the direct manipulating ZBTB16 via the shZBTB16 stereotaxic injection into the striatal regions. Via the RNA QauntSeq analysis, we first investigated the striatal transcriptomic changes by comparing the normal aging of wildtype (WT) mice and the pathological aging of amyloid precursor protein/presenilin 1 (APP/PS1) mice. Based on the transcriptomic analysis, we found that ZBTB16 could be a potent target gene of AD treatment and confirmed highly upregulated expression of the ZBTB16 in the striatal regions. Finally, we analyzed whether ZBTB16 is involved in the transcriptional regulation of the striatal transcriptome and deficits in behavior may be rescued by knockdown of the ZBTB16 expression.
Five-month-old, 6-month-old, and 12-month-old male APPswe/ PSEN1 dE9 transgenic mice and their littermate WT mice were used. Per cage, 4 or 5 mice were housed in group on a reversed 12-hour of day-night cycle at constant temperature and humidity. Water and food were supplied ad libitum. All experimental procedures for the usage and the treatment of animals were approved by the KIST Institutional Animal Care and Use Committee.
After habituation in a test chamber (45 cmأ—45 cmأ—45 cm) at day 1, mice were subjected to a familiarization in day 2 in the chamber with 2 identical red objects and were allowed to freely explore. After the familiarization, animals were placed in the chamber with one object replaced to a novel yellow object. The exploring times spent with each object were analyzed and calculated as a percentage of the total time of interaction.
Y-maze is a black maze apparatus with 3 arms (35 cmأ—10 cmأ—10 cm) angled 120آ° from each other. Mice were placed in the center of Yâ€گmaze and allowed to freely explore the maze for 10 minutes. The sequence of the arm entry to each arm was analyzed to analyze alternation percentage. Entry was counted as correct only when all 4 limbs were placed in the arm.
Three-chamber box, a Plexiglass box with 2 dividers each with openings (62 cmأ—41 cmأ—21.5 cm), was used. On the habituation, free explore was allowed for all 3 chambers for 10 minutes. In the test session, mice were placed in the middle, and 2 plastic objects and a stranger (S1) mouse placed into metal cages on each side. For the first session of sociability, mice were allowed to freely explore all chambers for 10 minutes. For the second session of social cognition, S1 was moved to the opposite chamber and additional stranger mouse (S2) was placed in the chamber that previously occupied by S1. The time spent and the number of entries into 3 chambers were analyzed.
Mice then underwent stereotaxic surgery (David Kopf Instruments, Tujunga, CA, USA) for the lentivirus-mediated delivery of the knockdown construct into the dorsal striatum. Mice were anesthetized with a cocktail of the ketamine (120 mg/kg) and the xylazine (6 mg/kg). Knockdown shRNA against ZBTB16 (TL507125, Origene, Rockville, MD, USA), shRNA scrambled control was packaged into pseudo-lentivirus particles following the manufacturer instructions (EndoFectin, EFL-1001-01, Genecopoeia, Rockville, MD, USA). Zero point five microliter of lentivirus was injected bilaterally using the Hamilton syringe with a 33G needle (WPI) at an injection rate of 0.05 خ¼L/min into the striatum (anteriorposterior +0.98, mediolateral آ±1.50, and dorsoventral -3.25, -3.5, -3.75 [from dura]) for each
injection locale of the striatum.
Blotted membranes were blocked using TBS-T with 5% bovine serum albumin solution and membranes were incubated with primary antibodies (mouse anti-ZBTB16 monoclonal antibody, 1:500, 39987, Active Motif, Carlsbad, CA, USA; mouse anti-خ² actin, 1:1,000, sc-47778, Santa Cruz Biotechnology, Inc., Dallas, TX, USA) overnight at 4آ°C. After the primary antibody treatment, secondary antibodies (donkey anti-mouse IgG-HRP, 1:2,000, sc-2318, Santa Cruz Biotechnology, Inc.) were treated in room temperature for 2 hours, followed by the visualization process with the chemiluminescent substrate of SuperSignal West Pico (Thermo Fisher Scientific Inc., Waltham, MA, USA). Chemiluminescent signals were visualized on the ImageQuant LAS4000 (GE Healthcare Life Sciences, Little Chalfont, UK) and ImageJ software (National Institute Health, Bethesda, MD, USA) was used to measure and analyze the densitometry analysis of the signal from bands.
RNA samples from the striatum were analyzed by quantitative real-time polymerase chain reaction (qPCR) experiments to confirm the expressional alteration of several differentially expressed genes (DEGs). Total RNA samples from the striatum were extracted and reverse transcribed via ReverTra Ace qPCR RT Master Mix (Toyobo, Osaka, Japan). RNA samples were analyzed by the reverse transcription qPCR with the appropriate primer sets using cDNA via THUNDERBIRD SYBR qPCR Master Mix (Toyobo).
Total RNA was isolated using Trizol reagent (Invitrogen, Waltham, MA, USA). RNA quality was assessed by Agilent 2100 bioanalyzer using the RNA 6000 Nano Chip (Agilent Technologies, Amstelveen, The Netherlands), and RNA quantification was performed using ND-2000 Spectrophotometer (Thermo Fisher Scientific Inc., Waltham, MA, USA). For each total RNA sample, the library construction was performed using QuantSeq 3آ´ mRNA-Seq Library Prep Kit (Lexogen GmbH, Wien, Austria) according to the manufacturer’s instructions. High-throughput sequencing was performed as single-end 75 sequencing using NextSeq 500 (Illumina, Inc., San Diego, CA, USA).
QuantSeq 3آ´ mRNA-Seq reads were aligned using Bowtie2. Bowtie2 indices were either generated from the genome assembly sequence or the representative transcript sequences aligning to the genome and transcriptome. The alignment file was used for assembling transcripts, estimating their abundances and detecting differential expression of genes. DEGs were determined based on the read counts and the fold change level from multiple alignments using coverage in bedtools.
Student unpaired t-test was used to compare data from WT group and APP/PS1 for noble object recognition (NOR) test, 3-chamber test (3CT), and the Western blot analysis. For the rest of results, 1-way analysis of variance (ANOVA) was used to analyze the effect of the APP/PS1 mutation and the lentivirus-mediated knockdown of shZBTB16 for the comparison of 3 groups in each analysis. Bonferroni multiple comparisons test method after the ANOVA analysis was used for the post hoc comparisons. P-values smaller than 0.05 were considered to be significantly different among comparisons from the statistical results (asterisks, *P <0.05, **P <0.01, ***P <0.001, ****P < 0.0001). Data were presented with bar graphs of the meanآ± standard error of the mean with error bars.
To identify significant gene expression changes in the striatal region associated with early stage of AD pathogenesis, 3آ´ mRNA-sequencing was performed with total RNA isolated from striatum region of 6-month-old and 12-month-old APP/PS1 mice and their littermate controls. DEG selection analysis was performed to select the potential target genes of AD pathogenesis (Fig. 1A). We identified early AD-specific 33 DEGs (Fig. 1B, C). To confirm the gene expression changes of the striatum in early stage AD, the expression level of the DEGs in the striatum of the 6-month-old APP/PS1 mice were analyzed by qPCR and we found that the expression levels of multiple genes were significantly altered (Fig. 1C). In previous study, we reported the functional role of MeCP2 in the striatum region in AD pathogenesis [5]. In matured neurons, MeCP2 is known to bind not only to methylated CpG (mCG) but also to non-CpG methlylated cytosines (mCH, H denotes A, C, or T) [9]. To select a single target for further analysis among selected DEGs, we compared mCG and mCH levels of the mouse striatal genome through the Brain Cell Methylation Viewer [10]. There was no apparent difference in mCG level among DEGs, but when the mCH level was analyzed, the level of methylation within the DNA was specifically increased in the gene body region of the ZBTB16 (Fig. 2). Based on this in-silico analysis, it was inferable that the change in MeCP2 expression and its subsequent epigenetic regulation could specifically affect the regulation of the ZBTB16 expression. Additionally, because pathway enrichment analysis results of the early striatal transcriptome showed major changes in pathways of the regulation of gene expression and protein homeostasis (data not shown), we focused on ZBTB16, which is being studied as a multifunction protein for both functions [4,11,12]. Among 33 DEGs, the increase of striatal ZBTB16 levels in 6-month-old APP/PS1 were additionally detected via the western blot analysis (Fig. 1D). Taken together, the analysis of the striatal transcriptome using 3’ mRNA-sequencing suggest that there may be noticeable changes in the gene expression profile including ZBTB16 throughout the early pathogenesis of AD and the manifestation of striatal dysfunction leading to the cognitive decline in the APP/PS1 mice.
In order to confirm the functional role of ZBTB16 and its increased level of expression in the striatum in the aspect of the transcription factor, an additional bioinformatic study was performed to analyze the upstream master regulator of the striatal transcriptomic alteration. Through this, we confirmed that ZBTB16 is one of the main transcription factors of the striatal transcriptome change, and also verified that multiple genes related with the brain function can be affected by the alteration of ZBTB16 expression. For the 560 genes of the early AD and 791 genes of the late AD defined in QuantSeq analysis, the chromatin immunoprecipitation-X enrichment analysis 3 (ChEA3) transcription factor enrichment analysis was performed using its database established by the integration of the previously curated database reports (Fig. 3A) [13]. Among transcription factors with a ChEA3 score of 1,000 or less in comparison set of the early AD and the late AD, ZBTB16 was ranked 123 out of the 1,234 transcription factors and 229 out of the 1,220 transcription factors, respectively. In addition, 53 genes in the early AD and 65 genes in the late AD were revealed as overlapping targets within the multiple databases and annotated as downstream target genes of ZBTB16. Numbers of target genes were also confirmed for each of the 5 integrated databases in the ChEA3 (Fig. 3B). Additional functional annotation of ZBTB16’s target genes also verified that the overlapping genes were involved in a number of neuronal function pathways including the neurogenesis and the neuron death pathway (Fig. 3C). Through this, it was confirmed that the expression changes of ZBTB16 could induce significant changes in the transcriptome of the striatal region.
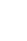
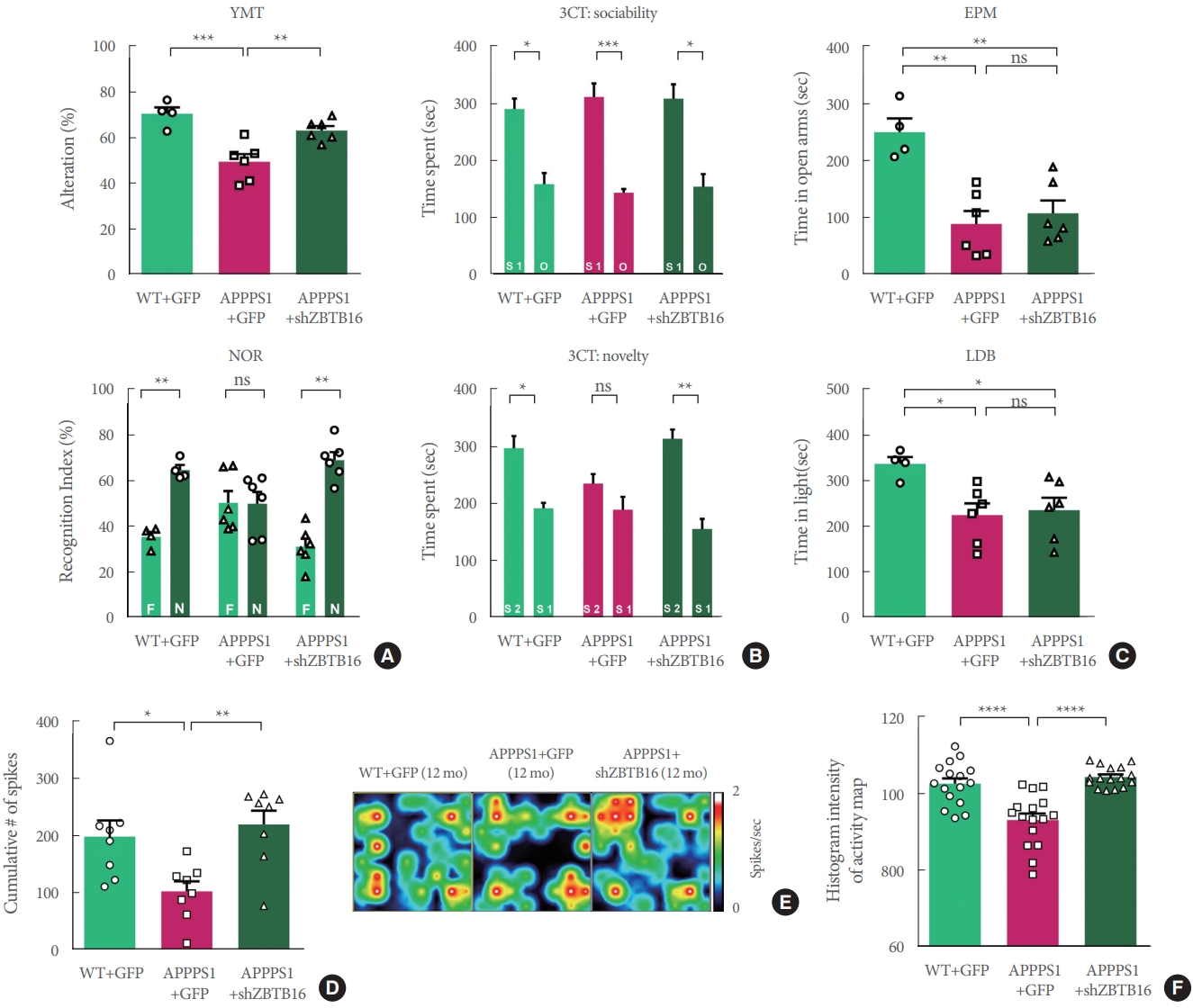
In order to confirm that the behavioral changes of 6-month-old APP/PS1 can be restored by ZBTB16 manipulation, a number of behavioral experiments testing cognitive function were additionally confirmed. The increased expression level of ZBTB16 was reduced to its normal level by using shZBTB16 and APP/PS1+GFP (green fluorescent protein) and APP/PS1 +shZBTB16 mouse showed normal locomotion (data not shown). Three behavioral experiments of Y-maze test (YMT), NOR test, and 3CT were performed to verify changes of the cognitive function in the APP/PS1+GFP and the APP/PS1+shZBTB16 mice. In YMT and NOR test, there was no difference between the 3 groups as previously characterized for the early stage of APP/PS1 (Fig. 4A). We confirmed 3CT-specific cognitive decline in the second session of the test. In addition, it was confirmed that this cognitive dysfunction in the 3CT was restored through knockdown of ZBTB16, confirming the improvement of social cognitive function through the restoration of striatal function (Fig. 4B). In addition, as a result of the anxiety level test verified through the elevated plus maze (EPM) test and the light-dark box (LDB) test, it was confirmed that the anxiety level increased in the APP/PS1+GFP was recovered through knockdown of ZBTB16 (Fig. 4C). Based on the behavioral analysis, it was verified that defective behavioral changes in the APP/PS1 could be restored through the normalization of the increased striatal ZBTB16 expression in the early stage of the APP/PS1 mice.
The same behavioral experiments were performed to confirm whether the knockdown effect at the 6 months could last until 12 months. As a result, in the age of 12 months, cognitive deficits in APP/PS1+GFP group were recovered in 3CT, YMT, and NOR test by the knockdown of ZBTB16 (Fig. 5A, B), but the manipulation did not recover the deficits in the EPM and LDB tests as shown in APP/PS +GFP and APP/PS1 +shZBTB16 mice (Fig. 5C). Also, when the striatal activity was analyzed using the multielectrode array (MEA), it was confirmed that the neural activity of the striatum, which was reduced in APPPS1 at 12 months of age, was restored by the knockdown of ZBTB16 (Fig. 5D-F). Overall, all the cognitive and behavioral dysfunctions except the anxiety level were recovered at 12 months of age by the effect of ZBTB16 knockdown at 6 months of age, and the rescue effect was also verified by the MEA analysis of the striatum activity.
The role of striatal ZBTB16 as a transcription factor in AD has not yet been clearly elucidated. And the involvement of ZBTB16 in the striatum undergoing pathological aging and the resulting behavioral phenotypes has not yet been explored. In this study, we found that the alteration in ZBTB16 that occurred in the early ages of the pathologically aging striatum coalesces with the expressional alteration of the target genes regulated by ZBTB16, while causing anxiety-like behavior, social memory deficits, and changes in neuronal activity. Furthermore, regulating the ZBTB16’s expression by knockdown may alleviate dysfunction throughout the early and late ages of APP/PS1. The early treatment of ZBTB16 knockdown in the striatum of APP/PS1 mice correlated with rescued cognition that continued into later age. This study demonstrates that perturbation of the gene expression during pathological aging may influence behavioral phenotypes and reveals a potent approach in targeting downstream genes of ZBTB16 in the striatum of AD patients.
In order to confirm the expression change of ZBTB16 as a target gene subject to epigenetic regulation of MeCP2, we performed in-silico analysis of the methylation level of the gene body of ZBTB16 and confirmed that DNA methylation of this region showed a significant difference compared to other DEGs. Prior to our report revealing the epigenome alteration of the striatal MeCP2 in AD mouse model [5], several reports discovered the regulatory networks between the DNA methylation and the recruitment of MeCP2 upon the gene body of ZBTB16 and their functional roles changing the expression level of ZBTB16 [6,7]. Our results were consistent with these reports, and through this, it is worthwhile to investigate the functional change of ZBTB16 by the MeCP2 regulation in terms of the rescue phenomenon of the cognitive impairment of APP/PS1.
ZBTB16 has been primarily studied as a transcriptional factor, binding directly to the target gene promoter region through its zinc fingers, or interacting with the histone deacetylase complex through the N-terminal BTB/POZ (Broad Complex, Tramtrack, bric-a-brac/Pox virus and zinc finger) domain. ZBTB16 mediated transcriptional repression by direct binding of ZBTB16 to target gene promoters involves genes associated with cell cycle progression and have pro-proliferative functions [14]. Representative target genes such as c-myc is a proto-oncogene that has various functions ranging from promoting cell proliferation to independently inhibiting cell differentiation [15]. In addition, cyclin A2 is another ZBTB16 targeted gene that is required for S-phase progression and preparation for mitotic entry, of which may be halted by cyclin A2 complex inhibition [16,17]. Interestingly, these genes that are involved in the cell cycle are closely tied to the neurodegenerative state, where amyloid beta oligomers may induce aberrant reentry into the cell cycle, causing neuronal death in familial AD animal models [18,19]. As a direct inhibitor of c-Myc and cyclin A2 expression, ZBTB16 may be distinguished as a potential modulator of the neuronal cycle and can block neuronal death. Therefore, it would be plausible to expect that the inhibitory effect of ZBTB16 on the cell cycle genes may be mitigated at the transcriptional or translational level. Furthermore, the increased ZBTB16 expression was sustained in the striatum of the APP/PS1 mice, suggesting that the role of ZBTB16 may vary according to AD-like pathological conditions, despite its potential role as a transcriptional regulator. Interestingly, although cell cycle event (CCE) markers show overall elevated levels in the brain of AD transgenic models, there is wide variability in CCE by brain region and transgenic models [20]. Therefore, results in our study indicate that the transcriptional inhibition regulating CCE in the context of AD pathology shows a different pattern and this may be an indication of cell cycle inhibition specific to the striatum.
At the same time, whether ZBTB16 functions either as a transcriptional repressor or autophagic regulator is largely dependent on the cellular localization of ZBTB16. In the cytoplasm, ZBTB16 itself is targeted for ubiquitination and degradation, or may act as an adaptor protein for the CUL3-ROC1 ubiquitination machinery, which implicates the multifaceted role of ZBTB16 both as a transcriptional and autophagic regulator. The ZBTB16-Cul3-Roc1 E3 ligase complex works to ubiquitinate ATG14L, leading to the degradation of ATG14L and ultimately halts the elongation phase of the autophagosome formation via phosphatidylinositol 3-kinase class III complex [21,22]. Previous studies have established that p62 directly binds with LC3 on autophagosomal membranes through the LC3 interacting region, and also binds to ubiqutinated proteins allowing for autophagy substrates to associate with the autophagosome [23-25]. Furthermore, LC3 is found to colocalize with ubiquitin in the brain in the event of disturbed autophagosome formation. To this end, we also the correlation of ZBTB16 levels with autophagic phenotype markers, LC3 and p62 in the striatum of APP/PS1 mice. Whereas ZBTB16 levels were consistently elevated through the early and late ages of APP/PS1 mice, p62 and LC3 showed abnormal patterns of expression at an early age, providing evidence of disturbed autophagy with the presence of AD-like pathology in the striatum. However, through a recent report, it was clarified that the mechanism of autophagy dysfunction in PS1 mutation is lysosome dysfunction among the 2 components at the stage of autophagosome and lysosome fusion [26]. Additionally, it has been suggested that ATG14L may function in autolysosome formation through homo-oligomerization [27]. Therefore, it can be recommended that strategies to improve the dysfunction of the autophagy process by the manipulation of ZBTB16 should be conducted in depth not only on autophagosome formation but also on the entire autophagy process.
NOTES
Funding/Grant Support
This work was supported by the Intramural grants by Korea Institute of Science and Technology (Grant No. 2E31522, 2E31700, 2E31524) to HII.
REFERENCES
1. Sweeney MD, Montagne A, Sagare AP, Nation DA, Schneider LS, Chui HC, et al. Vascular dysfunction-The disregarded partner of Alzheimer’s disease. Alzheimers Dement 2019;15:158-67. PMID: 30642436


2. Herrup K. The case for rejecting the amyloid cascade hypothesis. Nat Neurosci 2015;18:794-9. PMID: 26007212



3. Huguet G, Temel Y, Kadar E, Pol S, Casaca-Carreira J, Segura-Torres P, et al. Altered expression of dopaminergic cell fate regulating genes prior to manifestation of symptoms in a transgenic rat model of Huntington’s disease. Brain Res 2019;1712:101-8. PMID: 30711400


4. Usui N, Berto S, Konishi A, Kondo M, Konopka G, Matsuzaki H, et al. Zbtb16 regulates social cognitive behaviors and neocortical development. Transl Psychiatry 2021;11:242. PMID: 33895774




5. Lee S, Kim TK, Choi JE, Choi Y, You M, Ryu J, et al. Dysfunction of striatal MeCP2 is associated with cognitive decline in a mouse model of Alzheimer’s disease. Theranostics 2022;12:1404-18. PMID: 35154497



6. Khan AW, Ziemann M, Rafehi H, Maxwell S, Ciccotosto GD, ElOsta A. MeCP2 interacts with chromosomal microRNAs in brain. Epigenetics 2017;12:1028-37. PMID: 29412786



7. He Y, Hariharan M, Gorkin DU, Dickel DE, Luo C, Castanon RG, et al. Spatiotemporal DNA methylome dynamics of the developing mouse fetus. Nature 2020;583:752-9. PMID: 32728242




8. Puszyk W, Down T, Grimwade D, Chomienne C, Oakey RJ, Solomon E, et al. The epigenetic regulator PLZF represses L1 retrotransposition in germ and progenitor cells. EMBO J 2013;32:1941-52. PMID: 23727884



9. Chen L, Chen K, Lavery LA, Baker SA, Shaw CA, Li W, et al. MeCP2 binds to non-CG methylated DNA as neurons mature, influencing transcription and the timing of onset for Rett syndrome. Proc Natl Acad Sci U S A 2015;112:5509-14. PMID: 25870282



10. Liu H, Zhou J, Tian W, Luo C, Bartlett A, Aldridge A, et al. DNA methylation atlas of the mouse brain at single-cell resolution. Nature 2021;598:120-8. PMID: 34616061




11. Pennartz CM, Ito R, Verschure PF, Battaglia FP, Robbins TW. The hippocampal-striatal axis in learning, prediction and goal-directed behavior. Trends Neurosci 2011;34:548-59. PMID: 21889806


12. Abd-Elrahman KS, Hamilton A, Ferguson SSG. mGluR5 regulates ZBTB16 pathway of autophagy in Alzhimer’s disease in a sex-specific manner. Faseb Journal 2019;33(Suppl 1):810.5. 
13. Keenan AB, Torre D, Lachmann A, Leong AK, Wojciechowicz ML, Utti V, et al. ChEA3: transcription factor enrichment analysis by orthogonal omics integration. Nucleic Acids Res 2019;47:W212-24. PMID: 31114921




14. Liu TM, Lee EH, Lim B, Shyh-Chang N. Concise review: balancing stem cell self-renewal and differentiation with PLZF. Stem Cells 2016;34:277-87. PMID: 26676652



15. Pelengaris S, Khan M, Evan G. c-MYC: more than just a matter of life and death. Nat Rev Cancer 2002;2:764-76. PMID: 12360279



16. Gong D, Ferrell JE Jr. The roles of cyclin A2, B1, and B2 in early and late mitotic events. Mol Biol Cell 2010;21:3149-61. PMID: 20660152



17. Gopinathan L, Tan SL, Padmakumar VC, Coppola V, Tessarollo L, Kaldis P. Loss of Cdk2 and cyclin A2 impairs cell proliferation and tumorigenesis. Cancer Res 2014;74:3870-9. PMID: 24802190




18. Yang Y, Varvel NH, Lamb BT, Herrup K. Ectopic cell cycle events link human Alzheimer’s disease and amyloid precursor protein transgenic mouse models. J Neurosci 2006;26:775-84. PMID: 16421297



19. Varvel NH, Bhaskar K, Patil AR, Pimplikar SW, Herrup K, Lamb BT. Abeta oligomers induce neuronal cell cycle events in Alzheimer’s disease. J Neurosci 2008;28:10786-93. PMID: 18945886



20. Li L, Cheung T, Chen J, Herrup K. A comparative study of five mouse models of Alzheimer’s disease: cell cycle events reveal new insights into neurons at risk for death. Int J Alzheimers Dis 2011;2011:171464. PMID: 21912750




21. Zhang T, Dong K, Liang W, Xu D, Xia H, Geng J, et al. G-proteincoupled receptors regulate autophagy by ZBTB16-mediated ubiquitination and proteasomal degradation of Atg14L. Elife 2015;4:e06734. PMID: 25821988




22. Obara K, Ohsumi Y. Atg14: a key player in orchestrating autophagy. Int J Cell Biol 2011;2011:713435. PMID: 22013444




23. Pankiv S, Clausen TH, Lamark T, Brech A, Bruun JA, Outzen H, et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J Biol Chem 2007;282:24131-45. PMID: 17580304


24. Stolz A, Ernst A, Dikic I. Cargo recognition and trafficking in selective autophagy. Nat Cell Biol 2014;16:495-501. PMID: 24875736



25. Chun Y, Kim J. Autophagy: an essential degradation program for cellular homeostasis and life. Cells 2018;7:278. PMID: 30572663



26. Lie PPY, Yoo L, Goulbourne CN, Berg MJ, Stavrides P, Huo C, et al. Axonal transport of late endosomes and amphisomes is selectively modulated by local Ca(2+) efflux and disrupted by PSEN1 loss of function. Sci Adv 2022;8:eabj5716. PMID: 35486730



27. Liu R, Zhi X, Zhong Q. ATG14 controls SNARE-mediated autophagosome fusion with a lysosome. Autophagy 2015;11:847-9. PMID: 25945523



Fig.آ 1.
Quantitative gene expression profiling analysis of the striatal transcriptome of the amyloid precursor protein/presenilin 1 (APP/PS1) mice revealed striatum-specific differentially expressed genes (DEGs) including ZBTB16. (A) Summarized flowchart of the 3’ mRNA-sequencing analysis showing the criteria of DEG selection using the fold change (FC) values from the quantitative RNA sequencing data. (B) Heat-map presentation of the gene expression FC values of 33 DEGs from the mRNA-sequencing results from individual samples of wildtype (WT) and APP/PS1 mice. (C, D) Additional validation of gene expression of ZBTB16 in the mRNA level (C, quantitative polymerase chain reaction [qPCR]) and in the protein level (D, western blot analysis). AD, Alzheimer disease. *P<0.05. **P<0.01.
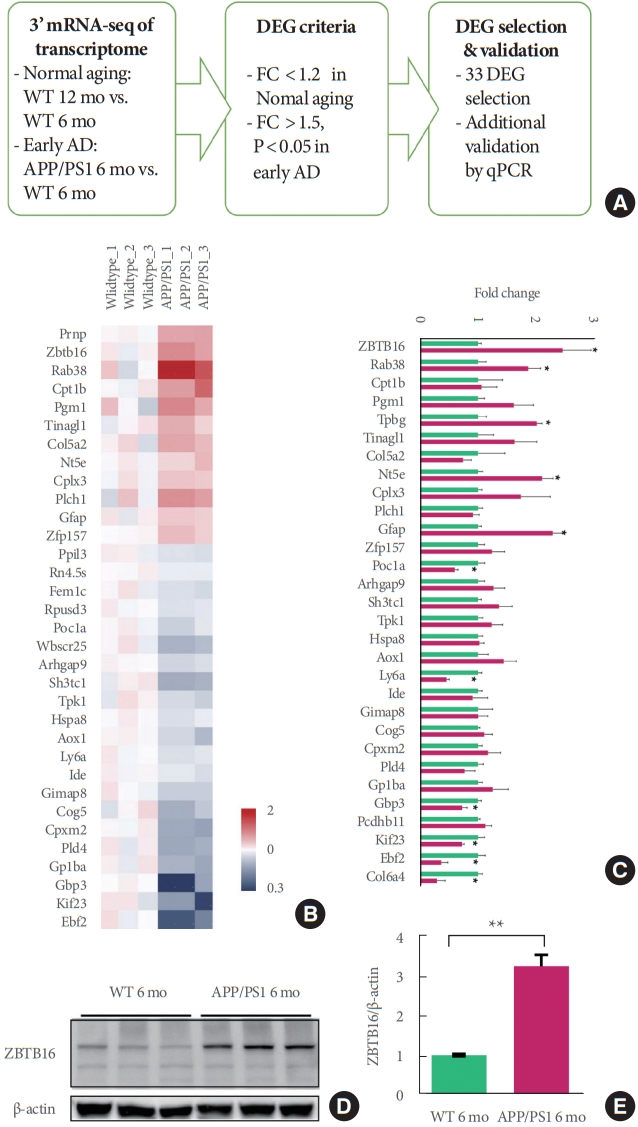
Fig.آ 2.
DNA methylation analysis with the resolution of single-cell via Brain Cell Methylation Viewer showed specific mCH (methylated non-CpG cytosine, H denotes A, C, or T) patterns of the ZBTB16 gene body. Single-cell methylome analysis via Brain Cell Methylation Viewer was used to compare the level of the mCH and the methylated CpG levels of 33 differentially expressed genes (DEGs). When confirming the gene body methylation level of the 12 increased DEGs of the early Alzheimer disease (bottom panels), the mCH level for the gene body of ZBTB16 was specifically increased (upper box). And this increase in mCH methylation was specific to D1 MSN and D2 MSN, which are the main neuron types of striatum.
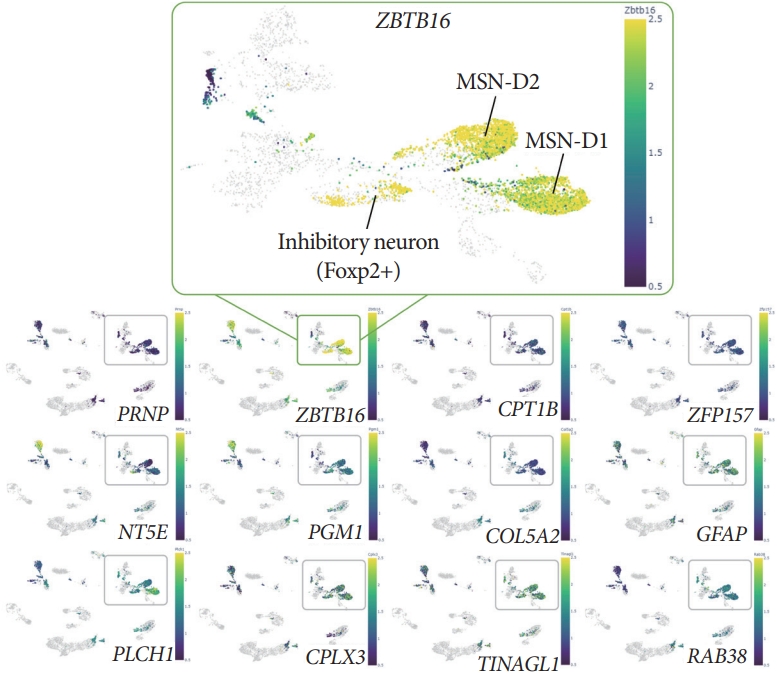
Fig.آ 3.
Transcriptomic alteration of the striatal region of amyloid precursor protein/presenilin 1 (APP/PS1) was associated with the function of zinc finger and BTB domain-containing protein 16 (ZBTB16) as a transcription factor. (A) Brief schematic flowchart of the chromatin immunoprecipitation-X enrichment analysis 3 (ChEA3) transcription factor enrichment analysis of the gene sets selected from the APP/PS1 striatal transcriptome changes in chapter 1. (B) Detail information of ZBTB16 from the ChEA3 transcription factor enrichment analysis for the altered transcriptome of the early Alzheimer disease (AD) and the late AD. Abbreviated 5 labels in the last column matched with the information of each library listed in panel A. (C) Functional annotation categories for the downstream overlapping genes of ZBTB16. WT, wildtype; DEG, differentially expressed genes; FC, fold change.
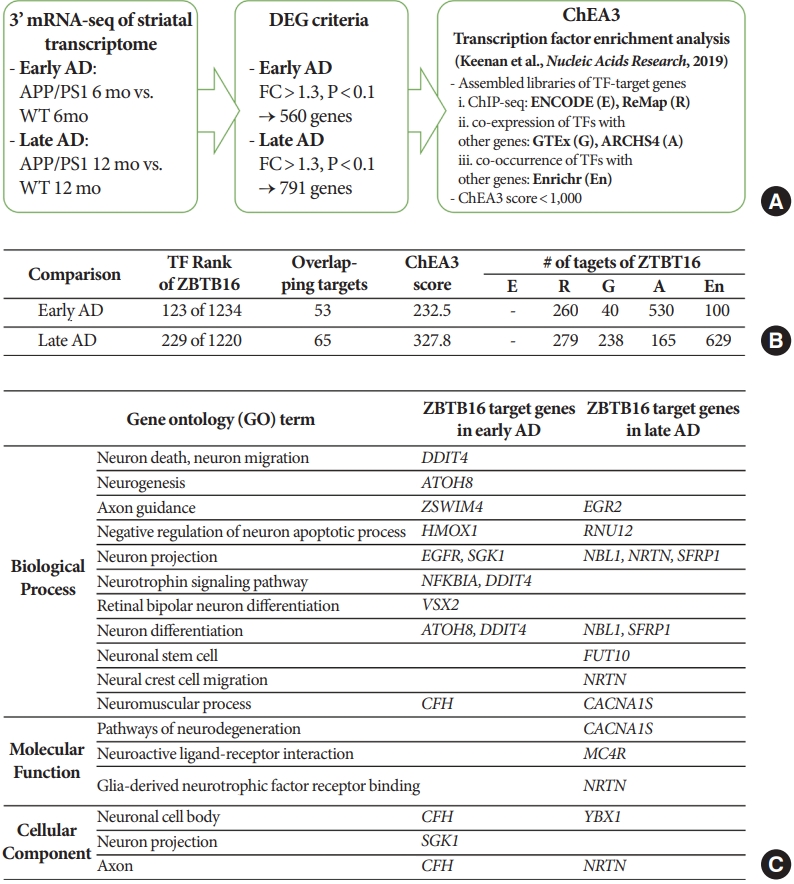
Fig.آ 4.
Behavioral defects were rescued by knockdown of striatal ZBTB16 of 6-month-old amyloid precursor protein/presenilin 1 (APP/PS1). (A) Cognitive function of wildtype (WT) and APP/PS1 mice of 6 months of age to validate the zinc finger and BTB domain-containing protein 16 (ZBTB16) knockdown effect. Memory function of WT+GFP (green fluorescent protein), APP/PS1+GFP, and APP/PS1+shZBTB16. Alterations in Y-maze test (YMT) and recognition index of noble object recognition (NOR) test are indicated by percentage. N and F indicate Novel and Familiar objects, respectively. (B) Time spent in chambers shown as a measure of sociability in the first session of the 3-chamber test (3CT) of WT+GFP, APP/PS1+GFP, and APP/PS1+shZBTB16 groups. And Time spent in chambers as a measure of novelty-seeking in the second session of 3CT. The early stage of APP/PS1 showed deficit in social novelty phenotype. And these deficits were rescued by ZBTB16 knockdown. S1 indicates stranger 1, S2 indicates stranger 2, and O indicates object. (C) In elevated plus maze (EPM) and light-dark box (LDB) test, time spent on the open arm and light box were shown in seconds. ns, not significant. *P<0.05. **P<0.01. ***P<0.001.
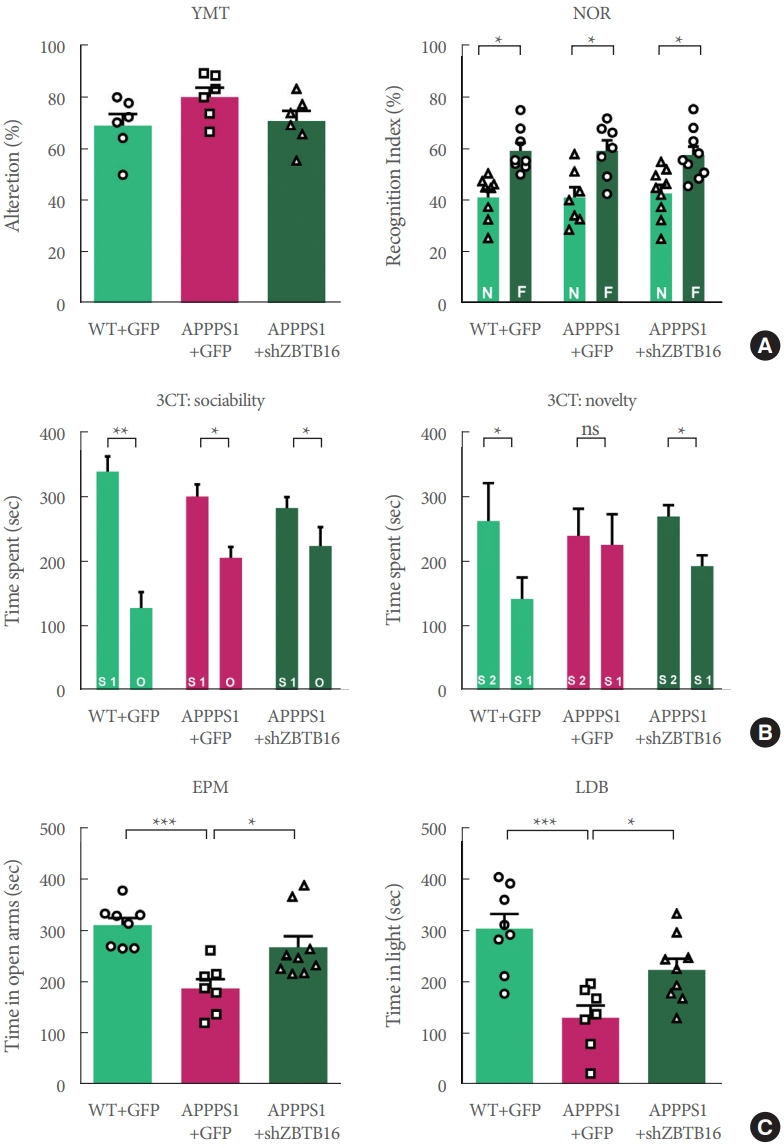
Fig.آ 5.
Knockdown of zinc finger and BTB domain-containing protein 16 (ZBTB16) at 5 months rescued behavioral defects in 12-month-old amyloid precursor protein/presenilin 1 (APP/PS1). (A) Similar rescue patterns in 3-chamber test (3CT) were also observed in the 12-month-old APP/PS1+shZBTB16 group (social novelty). (B) In Y-maze test (YMT) and noble object recognition (NOR) test, cognitive impairments in APP/PS1 (12-month-old) were also rescued by ZBTB16 knockdown. (C) Deficits in anxietylike phenotypes of APP/PS1+GFP (green fluorescent protein) were not rescued by ZBTB16 knockdown in the elevated plus maze (EPM) and light-dark box (LDB) tests. (D) Bar graphs showing the cumulative number of firings from all stimulation currents (from 125 nA to 1,000 nA currents). (E, F) Representative heat-map image and quantification (histogram densitometry) bar graph of neuronal activity on the multielectrode array probes. APP/PS1+ZBTB16 knockdown group showed rescued patterns. ns, not significant. *P<0.05. **P<0.01. ***P<0.001. ****P<0.0001.