|
 |


- Search
| Int Neurourol J > Volume 18(3); 2014 > Article |
|
ABSTRACT
Interstitial cystitis (IC), also known as painful bladder syndrome or bladder pain syndrome, is a chronic lower urinary tract syndrome characterized by pelvic pain, urinary urgency, and increased urinary frequency in the absence of bacterial infection or identifiable clinicopathology. IC can lead to long-term adverse effects on the patient's quality of life. Therefore, early diagnosis and better understanding of the mechanisms underlying IC are needed. Metabolomic studies of biofluids have become a powerful method for assessing disease mechanisms and biomarker discovery, which potentially address these important clinical needs. However, limited intensive metabolic profiles have been elucidated in IC. The article is a short review on metabolomic analyses that provide a unique fingerprint of IC with a focus on its use in determining a potential diagnostic biomarker associated with symptoms, a response predictor of therapy, and a prognostic marker.
The main goal of this article is to provide the reader with an up-to-date summary of the main metabolic variations taking place in biofluids with respect to interstitial cystitis (IC)/pelvic bladder syndrome/bladder pain syndrome as well as of the analytical strategies employed to unveil urinary metabolomic biomarkers.
Metabolomics provides a global chemical fingerprint of the metabolism of cells and indicates physiological and pathological states of biological samples. Thus, the power of metabolomics allows unparalleled opportunity to query the molecular mechanisms of the disease. Metabolites are not merely the end products of gene/protein expression; rather, they are the result of the interaction of the genome with its environment in the cells [1]. Thus, analyzing metabolic differences between pathological and normal conditions could provide scientific insight into the underlying disease pathology.
Current metabolomics technologies may provide a comprehensive understanding of IC beyond the classical criteria, which are dependent on subjective observations. IC metabolomics provide the characteristics of the disease state and identify novel approaches to reduce clinical symptoms, including chronic bladder pain, a hallmark symptom of patients with IC. Despite the large number of review articles on metabolomics and its application in cancer and diabetes research [2,3], there is no comprehensive review on metabolite profiling to study disease mechanisms or biomarkers in IC.
Based on these current circumstances, we have organized this review article as follows. First, we present a short review on IC. Second, we discuss analytical techniques that are used in metabolomics, including computational methods for data processing. Next, we present an overview of the studies that have used metabolomics to reveal IC metabolic fingerprints. Finally, we discuss the future research directions and summarize our findings.
IC is characterized as a chronic bladder condition in which the disorder of the urinary bladder manifests as long-term symptoms that include urinary frequency/urgency or pain, pressure, discomfort, and nocturia. Chronic bladder pain is considered the hallmark symptom of patients with IC. Approximately 1 out of 77 people in the United States have been diagnosed with IC, and approximately 80% of patients are women [4,5]. IC has a long-lasting and adverse impact on the quality of life of patients, because IC suppresses physical function and social activity [4,5,6]. Its symptoms may be mild or severe and occasional or constant [7]. Many patients find that symptoms worsened with stress (either physical or mental stress). The medical costs associated with IC has increased and is currently estimated to exceed $100 million/yr in USA [8].
Unfortunately, etiologies of IC are largely unknown. No effective treatment or prescription of medications has been clearly suggested. Additionally, the methods for accurate diagnosis of this condition have not been established thus far. Thus, physicians are dependent on the inclusive and exclusive clinical diagnostic criteria based on National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) guidelines (e.g., evaluation and treatment based on results from urinalysis, urine culture, cystoscopy, biopsy of the bladder wall and urethra, and bladder distention) [7]. The evaluation of the patient's medical history, physical exam, and urine tests are needed for ruling out other conditions that might have caused similar symptoms.
For IC diagnosis, the process from the first doctor's visit to a definitive diagnosis in the urology clinic takes up a few years. The significant overlap between IC and other benign bladder dysfunction with similar urinary symptoms, including overactive bladder, provides another challenge for IC diagnosis. A standard diagnostic test specific to IC has not yet been introduced to the current clinical practice, and diagnosis is still dependent on subjective criteria. Thus, the identification of biomarkers of IC and the elucidation of the pathogenic mechanisms of biomarkers are clinical needs whose solution may lead to the discovery of new therapeutic approaches and biomarkers for better diagnosis of IC.
Efficient treatment regimens for IC have been another challenge to patients and urologists, which is also associated with the difficulty in diagnosis. Although there are multiple hypotheses about the primary cause of IC, the underlying pathologic mechanism remains completely undefined, which leads to delays in identifying effective therapeutic modalities. At this time, the Food and Drug Administration has approved only two treatments for IC. One is oral pentosan polysulfate. The other treatment is to place dimethyl sulfoxide into the bladder through a catheter. Another approach is to include procedures, such as hydrodistention, and oral pharmaceutical drugs, such as antihistamines, tricyclic antidepressants, and immune modulators. In general, symptom relief achieved with any of these strategies has been reported in only a small percentage of patients, and recurrence typically is evident within a few months [4,9]. For example, antibiotics and pentosan polysulfate have been suggested to relieve the bladder pain or discomfort associated with IC; however, this treatment is only effective in relieving bladder pain in about 30%-60% of patients and it accompanies unfavorable side effects, including nausea, diarrhea, gastric distress, and hair loss. Moreover, while the mechanism of action of pentosan polysulfate in IC has been uncovered, the drug has anticoagulant and fibrinolytic effects.
Several abnormalities have been reported in epithelial, endothelial, smooth muscle (detrusor), neuronal, or immune cells in IC patients. Bladder epithelium thinning, erosion, and increased infiltration of mast cells are some characteristics of IC histology, which suggest that the disease is mediated by a defect in the regenerative potential of bladder epithelial cells and/or by immune impairment [4,5]. Furthermore, urothelium derived from IC patients exhibited suppressed cell proliferation and increased cell permeability [10,11,12], and these findings are supported at the molecular level in that the expression of proproliferative proteins (e.g., cyclin D1) and barrier tight junction proteins (e.g., zonula occludens-1, occludin, and claudin 1, 4, and 8) is dysregulated [11,12]. Bladder endothelium of IC often displays an increased level of platelet-derived endothelial cell growth factor/thymidine phosphorylase, vascular endothelial growth factor, and substance P expression [13,14,15] compared to those of controls. Notably, the IC bladder epithelial cells exhibited intrinsic alterations in differentiation, neurotransmitter release, and potassium channel activity with elevated nitric oxide production, nuclear factor-κB activation, nerve fiber number, serum C-reactive protein [16], and neuropeptide Y and nerve growth factor production [16,17,18], suggesting the neural control of the urinary bladder [19].
Attempts to identify IC diagnostic markers have employed urine cytology, cystoscopy with and without hydrodistention and/or bladder biopsy, and biofluid-based assays [7,20,21,22]. However, cytology is nondiagnostic. Cystoscopic observation is invasive, expensive, and painful. Cystoscopy cannot distinguish patients from normal since cystoscopic appearance is normal in the majority of IC patients [21]. The potassium sensitivity test is also used for diagnosis. It is based on the response of whether IC patients are more sensitive to the potassium solution than to water when placed in the bladder. However, the potassium test is painful and not specific to IC diagnosis [22]. Urodynamics evaluation is another test; however, it is not necessary in all cases.
Metabolomics based on biofluids (including urine, plasma, saliva, fecal extract, and sputum et al.) have often been used to discover a noninvasive or minimally invasive biomarker test for other diseases, including cancer and diabetes [23]. In particular, assays of urine components are noninvasive and can be easily repeated, even by the patient. As one of many examples of urinary IC biomarkers, a small glycosylated peptide, called an antiproliferative factor (APF), was originally identified and characterized by Keay et al. [24] This short sialoglycopeptide consisting of 8 amino acids is 100% homologous to the putative sixth transmembrane domain of frizzled 8 (a Wnt receptor). Increased APF bioactivity is observed in urine specimens of IC patients. The measurement of APF in urine could segregate IC patients with 94% sensitivity and 95% specificity [25,26,27]. From in vitro systems, many reports, including those from our laboratory, demonstrated that APF regulates important signaling pathways, such as Akt, MAPK, β-catenin, or p53 signaling, leading to the suppression of proliferation of bladder epithelial cells and the increase in transurothelial cellular permeability [24,28,29,30]. The global and unbiased APF signaling network was uncovered using a state-of-the-art quantitative proteomics method and a systems biology approach [28,30,31,32,33].
Metabolomic profiling, or metabolomics, is the systemic study of the unique small chemical fingerprints in a biological sample. The collection of small-molecule profiles represents the end products of cellular processes in biological systems (e.g., cells, tissues, or organs). A small amount of biofluid (urine, plasma, etc.) [23] allows for the characterization of hundreds of metabolites that provides a functional readout of the metabolic state. Based on the accumulated knowledge-based database, metabolite profiles can provide the biological interpretation of metabolic perturbations unique in IC patients [34]. In this section, we review the current metabolomics techniques and analytical tools/software for IC metabolic profiling.
Metabolomic studies typically begin with sample collection followed by sample analysis. A number of analytical techniques are currently used for metabolomic studies depending on the particular metabolite of interest. Nuclear magnetic resonance spectroscopy (NMR, in most cases 1H-NMR), gas chromatography-mass spectrometry (GC-MS), and liquid chromatography-mass spectrometry (LC-MS) metabolic techniques are widely used methods of analysis [35]. Much of the original metabolomics analyses were performed using NMR spectroscopy. Currently, chromatographic techniques, such as GC and LC coupled with MS (GS-MS and LC-MS), are used more. Fourier transform spectrometry [36] and capillary electrophoresis-MS [37,38] are also the major spectroscopic techniques used in metabolomic analysis.
(1) NMR-based metabolomics analysis is robust and straightforward to implement. In particular, the reproducibility of NMR spectra is excellent [35], and NMR allows the easy identification of metabolites from 1-dimensional spectra. NMR is the best analytical tool for urine and plasma analysis due to its nondestructive nature, minimum sample requirement, quantitative ability, and reliable metabolite identification that provides detailed information on the structure. However, NMR suffers from a lower sensitivity than MS [39], and compound identification is less straightforward compared to MS methods.
(2) MS-based metabolomics is more sensitive than NMR-based metabolomics; however, it provides a higher risk of generating technical artifacts and measurement errors. MS is most often coupled to additional gas- or liquid-phase chromatography separation steps that allow for the stratification of complex mixtures. MS-based methods provide an overall large number of molecules that can be quantified. While sample preparation procedure for GC-MS, which includes extraction, drying, and concentration, is time-consuming, GC-MS is highly sensitive, reproducible, and cost-efficient. LC-MS also requires less comprehensive sample preparation than GC-MS.
There are advantages and limitations in the different metabolomic methods. In general, MS involves the use of more complex sample extraction methods, whereas NMR needs greater sample volumes. Both methods can provide identification of metabolites when their identity is initially unknown. These unknowns may eventually be identified using additional analytical methods, and thereby, can provide new insights into the phenotype or the disease [40]. Hence, a combination of different analytical techniques provides more information than a single tool when analyzing the complete metabolome.
Depending on the goal of the study, targeted or nontargeted metabolomics approaches may be selected. Targeted approaches focus on only a specifically defined subset of metabolites. Targeted approaches are typically used to quantify the measured metabolites on an absolute scale by using external or internal standards as a reference, whereas nontargeted approaches provide only semiquantitative measures, such as ion counts or areas under a curve using arbitrary units. Targeted methods with absolute quantification have the advantage in that measurements from different studies can easily be combined and compared. Targeted methods have lower technical variance since more time can be spent on measuring a smaller set of metabolic features. For the discovery of new processes and pathways, the use of nontargeted methods is recommended. By providing the widest possible range of metabolites, nontargeted metabolomics approaches can reveal global profiles to discover new molecules that are associated with the phenotype under investigation.
To identify the metabolites that are differentially expressed between the samples to possibly lead to biomarker selection, the large amount of data generated by this analysis should then be statistically processed. The key to identifying potential biomarkers is based on the difference in metabolite levels between the biological samples taken from the IC patients and normal (control) subjects.
Several statistical tools are currently used to analyze NMR and MS-based metabolomics datasets, which include XCMS, MZmine, MetAlign, MathDAMP, and LCMStats. For NMR data analysis, the significantly altered NMR peaks can be identified by using a probabilistic approach called Bayesian Quantification (BQuant) [39], which will allow the identification and quantification of metabolites in local regions of the 1H-NMR spectra. For preprocessing of data from MS-based metabolomics data, XCMS software, a web-based integrated metabolomics analysis platform for the determination of metabolic profile differences and metabolite identification can be applied to reading, detection, and alignment. MetaboSearch software [41] could be used to perform mass-based metabolite identification simultaneously against the four major metabolite databases: the Human Metabolome DataBase (HMDB), Madison Metabolomics Consortium Database, METLIN (a metabolomics database), and LipidMaps. The ultimate output would reveal an extracted metabolite profile. This can then be exported into software, such as MATLAB, for multivariate analysis to select metabolites that can further subdivide the IC from the controls [34].
To elucidate the patterns and connections of the identified metabolites, a number of statistical algorithms for metabolomic data analysis could be applied both in a supervised and unsupervised manner. The unsupervised methods that have been extensively used in metabolomic analysis are principal component analysis, hierarchical clustering, and self-organizing maps. Supervised methods include analysis of variance, partial least squares (PLS), hierarchical PLS, k-nearest neighbors, and discriminant function analysis. Using these methods samples can be clustered based on their metabolic fingerprints. Data integration and analysis is an important component of metabolomic studies, because a large amount of data is generated that is similar to proteomic and transcriptomic studies. The following databases could be used: HMDB (http://www.hmdb.ca/), METLIN (http://metlin.scripps.edu/), MassBank (http://www.massbank.jp), PubChem (http://ncbi.nim.nih.gov/), and Kyoto Encyclopedia of Genes and Genomes (http://www.kegg.jp/). To extract IC-associated biological features and to identify the altered metabolic pathways/networks in IC, intensive computational analyses for data integration with proteomics and genomics databases should be considered as described in our previous review [42].
In summary, given that metabolomics also has potential utility in several fields of other diseases, including disease prognosis, diagnosis, and drug discovery and development, metabolic signatures may serve as an alternative strategy for personalized therapy for IC patients with different phenotypes. The field of metabolomics may offer practical solutions to the challenges in IC research. Further research of the physiological change and mechanisms of IC requires more clinical data and powerful analytical strategies, especially structural identification techniques.
As mentioned above, the best diagnostic would include a noninvasive biomarker(s) test with high sensitivity and specificity. A non- or minimally invasive diagnostic method using biofluids (e.g., urine, blood, saliva, fecal extract, and sputum specimens) may play a significant role in IC with regard to early detection, diagnosis, prognosis, drug development, and sensitivity prediction to clinical treatments.
The most attractive biofluids in metabolomics are serum and urine [43,44,45]. Serum is a readily accessible and informative biofluid, making it ideal for early detection of a wide range of diseases. Monitoring of serum metabolites has several advantages due to its stability and less dilution effect [46]. Metabolite profiles of serum can be regarded as important indicators of physiological and pathological states and may aid in the understanding of the mechanism behind disease occurrence and progression on the metabolic level.
Urine is also an ideal bio-medium to study disease, because it is readily obtained and available with no required preparation by the patient, and it is less complex than other body fluids [47]. The ease of collection allows for serial sampling to monitor disease and therapeutic response. In terms of proteins and the prevalence of different cell types in blood, urine is much less complex, but nevertheless can contain disease biomarkers [4,6,23,48], specifically, excreted metabolites. Body fluids that are most proximal to a disease site can often provide a source of informative biomarkers [35,46,49]; therefore, urine-based diagnosis for IC is the most attractive strategy among other biofluids-based methods. However, analysis of urine samples poses several analytical challenges for metabolic profiling owing to wide variations in the ionic strength, pH, and osmolarity, particularly under conditions of physiological stress. Furthermore, urine samples typically have a huge dynamic range of metabolite concentrations and are subject to unpredictable dilution effects. Thus, to minimize artifacts arising from irrelevant biological variations, data can be normalized to creatinine concentrations, preferably by correcting volumes used for metabolomics assays instead of just correcting acquired data. Although creatinine is the best possible internal standard for correcting urine volume effects, creatinine levels can vary due to dietary intake and pathological conditions. Computational approaches for data normalization methods can be applied to reduce artifacts due to sample variability using currently developed probabilistic quotient- and median-fold changes in normalization strategies [50,51].
With the successful discovery of APF in IC patients' urine [10,24], urine analysis has already had some success in studies of this condition. Metabolomics has demonstrated promise in the development of diagnostic tools for IC. Previously, a few attempts to use metabolomics analysis to identify an IC signature have been published.
In 2003, Van et al. [52] has suggested that MS and NMR spectral patterns can segregate patients with IC from non-IC patients, including bacterial cystitis (40 females and 10 males). Asymptomatic controls included individuals that were age-, race-, and gender-matched to the IC patients (40 females and 10 males). Their NMR-based metabolomics analysis has identified 1,4-methyl-imidazole-acetic-acid and norepinephrine as potential diagnostic markers for IC. More recently, to distinguish the metabolites between patients with IC (n=10), those with bacterial cystitis (n=10), and healthy volunteers (n=10), Fukui et al. [53] has performed a nontargeted analysis using ultraperformance liquid chromatography-time-of-flight/mass spectrometry-based metabolomics. They used 5 IC patients and 5 healthy volunteers as a validation cohort. Their findings suggest that the urinary ratio of phenylacetylglutamine to creatinine can be correlated to the clinical grade of IC (e.g., IC patients with mild (I/II) to severe (III) symptoms when compared to healthy volunteers). However, the mechanism of why urine specimens from IC patients have an increased phenylacetylglutamine to creatinine is not well understood. Additionally, this finding, while promising, is based on a small number of patients. A larger, prospective cohort-based validation is needed to translate this ratio to a clinical setting, which may help elucidate the mechanism governing IC initiation and progression.
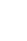
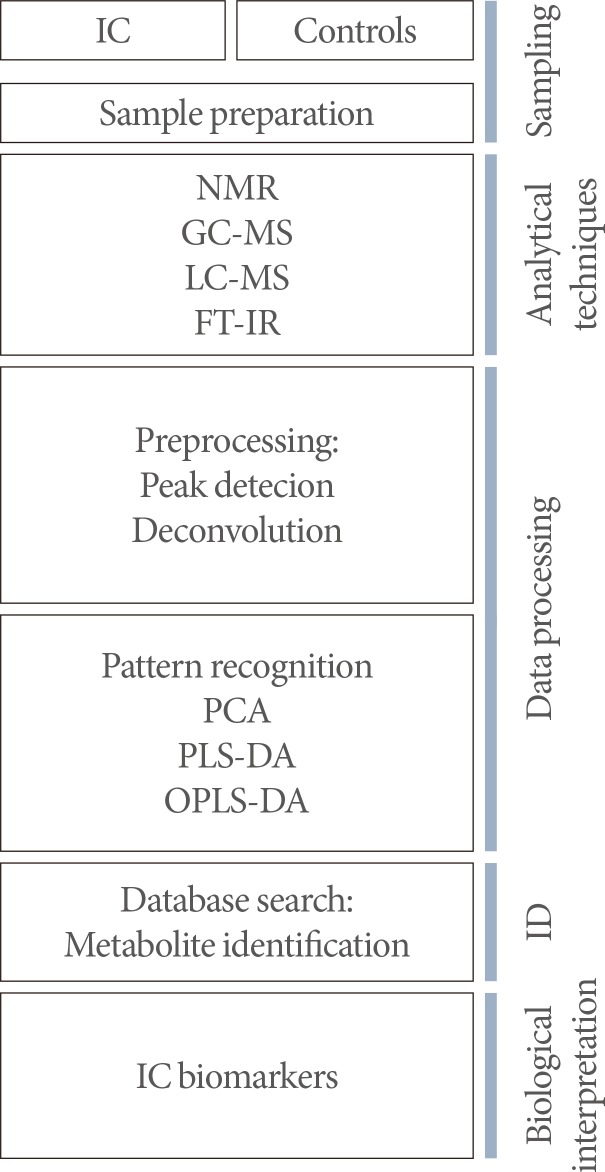
Collectively, a workflow for IC metabolic profiling is suggested in Fig. 1. Researchers are also aware that the ability to obtain a high quality sample along with sample collection, storage, and analysis are important factors that have large consequences on metabolic results. This fact underscores the need for standardized protocols. Recently the National Institutes of Health (NIH)/NIDDK established the "Multidisciplinary Approach to the Study of Chronic Pelvic Pain" (MAPP) Research Network to better understand the pathophysiology of urologic chronic pelvic pain syndromes (including IC and chronic prostatitis/chronic pelvic pain syndrome) and to provide a translational foundation for the future clinical intervention and an improved clinical management. This ambitious NIH-funded, nationally wide project has collected a multisite and longitudinal biorepository of human specimens derived from men and women with IC as well as those from the age and gender-matched healthy controls using a standardized protocol through the phase I MAPP Network [54,55].
As a future direction, we may consider several studies that should soon be answered by metabolomics approaches. To identify metabolic signatures as predictive biomarkers of disease progression and its related comorbidities, metabolome assessments need to be studied prior to disease onset and compared to metabolic changes during disease development. In particular, prospective studies can provide valuable insights into the mechanisms and processes altered during IC development and its comorbidities. Thus, this approach might be critical for biomarker discovery for IC. Currently, the most frequently applied drug and treatment for IC is pentosan polysulfate and hydrodistention, respectively. However, there are frequent recurrences with unfavorable side effects. Sensitivity prediction is a challenging task in the treatment of IC patients. Some patients with IC do not respond well to the current clinical therapies, and in some cases, therapies may cause severe toxicity and functional impairment. Hence, it is crucial to phenotype individual patients with high responsiveness to treatment and to recommend personalized therapy only for the selected patients. Altered metabolic profiles may be related to treatment response, and the perturbation pattern indicates functionally relevant perturbations in metabolism. The metabolite signature can also be used to identify mechanistic markers for drug or treatment response, which can then be used for screening and efficacy tests. For example, for a metabolic approach for responsiveness prediction to therapies, bio-specimens (such as urine or serum) could be used to derive metabolic profiles using NMR or MS-based metabolomics analysis. From the metabolic data, a proposed predictive model could be developed with high accuracy and sensitivity for metabolites in IC, which can serve as a crucial modulator of IC responsiveness to clinical therapies. The prospective studies should attempt to recruit patients who are newly diagnosed with IC without previous treatment. This longitudinal sample collection for metabolic analysis should be thoughtfully collected under the standardized clinical protocol. Given that it is important to diagnose IC at an early stage to initiate appropriate therapy and improve the patient's quality of life, future research need to consider the following: (1) prospective studies to identify IC biomarkers based on metabolite profiles; (2) dynamic metabolomics analysis to determine the metabolites associated with symptom progression; and (3) metabolome perturbation and drug/treatment response (the identification of drug side effects and/or functional biomarkers for drug action).
IC, a debilitating chronic bladder disease, significantly reduces quality of life of women and men. Diagnostic and treatment modalities, even subjective diagnostic tools, are largely unavailable. Metabolomics analysis combined with computational approaches could provide a fingerprint of the metabolism of the disease and the essential molecular details about regulatory mechanisms. Metabolomics will lead to new insights into the perturbations leading to IC and other essential information for the goal of improving diagnostic capability and treatment strategies for patients.
National Institutes of Health1R01DK100974-01U24 DK097154NIH NCATS UCLA CTSI UL1TR000124
Interstitial Cystitis Association
New York Academy of Medicine
Boston Children's Hospital
Interstitial Cystitis Association
New York Academy of Medicine
Boston Children's Hospital
ACKNOWLEDGEMENTS
This research was supported by: NIH grants 1R01DK100974-01, U24 DK097154, NIH NCATS UCLA CTSI UL1TR000124, IMAGINE NO IC Research Grant, the Steven Spielberg Discovery Fund in Prostate Cancer Research Career Development Award, Interstitial Cystitis Association (ICA) Pilot Grant, and a Fishbein Family IC Research Grant by ICA; New York Academy of Medicine; Boston Children's Hospital Faculty Development (to J.K.).
REFERENCES
1. Cho K, Mahieu NG, Johnson SL, Patti GJ. After the feature presentation: technologies bridging untargeted metabolomics and biology. Curr Opin Biotechnol 2014;28:143-148. PMID: 24816495.



3. Nair M, Sandhu SS, Sharma AK. Prognostic and predictive biomarkers in cancer. Curr Cancer Drug Targets 2014;14:477-504.


4. Persu C, Cauni V, Gutue S, Blaj I, Jinga V, Geavlete P. From interstitial cystitis to chronic pelvic pain. J Med Life 2010;3:167-174. PMID: 20968203.


5. Phatak S, Foster HE Jr. The management of interstitial cystitis: an update. Nat Clin Pract Urol 2006;3:45-53. PMID: 16474494.


6. Evans RJ, Sant GR. Current diagnosis of interstitial cystitis: an evolving paradigm. Urology 2007;69(4 Suppl):64-72. PMID: 17462483.


7. Hanno PM, Burks DA, Clemens JQ, Dmochowski RR, Erickson D, Fitzgerald MP, et al. AUA guideline for the diagnosis and treatment of interstitial cystitis/bladder pain syndrome. J Urol 2011;185:2162-2170. PMID: 21497847.


8. Anger JT, Zabihi N, Clemens JQ, Payne CK, Saigal CS, Rodriguez LV. Treatment choice, duration, and cost in patients with interstitial cystitis and painful bladder syndrome. Int Urogynecol J 2011;22:395-400. PMID: 20811877.



9. Hanno PM. Analysis of long-term Elmiron therapy for interstitial cystitis. Urology 1997;49(5A Suppl):93-99. PMID: 9146008.


10. Keay S, Zhang CO, Shoenfelt JL, Chai TC. Decreased in vitro proliferation of bladder epithelial cells from patients with interstitial cystitis. Urology 2003;61:1278-1284. PMID: 12809929.


11. Slobodov G, Feloney M, Gran C, Kyker KD, Hurst RE, Culkin DJ. Abnormal expression of molecular markers for bladder impermeability and differentiation in the urothelium of patients with interstitial cystitis. J Urol 2004;171:1554-1558. PMID: 15017219.


12. Zhang CO, Wang JY, Koch KR, Keay S. Regulation of tight junction proteins and bladder epithelial paracellular permeability by an antiproliferative factor from patients with interstitial cystitis. J Urol 2005;174:2382-2387. PMID: 16280852.


13. Kiuchi H, Tsujimura A, Takao T, Yamamoto K, Nakayama J, Miyagawa Y, et al. Increased vascular endothelial growth factor expression in patients with bladder pain syndrome/interstitial cystitis: its association with pain severity and glomerulations. BJU Int 2009;104:826-831. PMID: 19298410.


14. Tamaki M, Saito R, Ogawa O, Yoshimura N, Ueda T. Possible mechanisms inducing glomerulations in interstitial cystitis: relationship between endoscopic findings and expression of angiogenic growth factors. J Urol 2004;172:945-948. PMID: 15311005.


15. Li J, Micevych P, McDonald J, Rapkin A, Chaban V. Inflammation in the uterus induces phosphorylated extracellular signal-regulated kinase and substance P immunoreactivity in dorsal root ganglia neurons innervating both uterus and colon in rats. J Neurosci Res 2008;86:2746-2752. PMID: 18478547.



16. Chung SD, Liu HT, Lin H, Kuo HC. Elevation of serum c-reactive protein in patients with OAB and IC/BPS implies chronic inflammation in the urinary bladder. Neurourol Urodyn 2011;30:417-420. PMID: 21284020.


17. Ochodnický P, Cruz CD, Yoshimura N, Michel MC. Nerve growth factor in bladder dysfunction: contributing factor, biomarker, and therapeutic target. Neurourol Urodyn 2011;30:1227-1241. PMID: 21520250.


18. Liu HT, Chen CY, Kuo HC. Urinary nerve growth factor levels in overactive bladder syndrome and lower urinary tract disorders. J Formos Med Assoc 2010;109:862-878. PMID: 21195884.


19. Kilpatrick LA, Kutch JJ, Tillisch K, Naliboff BD, Labus JS, Jiang Z, et al. Alterations in resting state oscillations and connectivity in sensory and motor networks in women with interstitial cystitis/painful bladder syndrome. J Urol 2014;192:947-955. PMID: 24681331.



20. Johansson SL, Fall M. Clinical features and spectrum of light microscopic changes in interstitial cystitis. J Urol 1990;143:1118-1124. PMID: 2342171.


21. Ottem DP, Teichman JM. What is the value of cystoscopy with hydrodistension for interstitial cystitis? Urology 2005;66:494-499. PMID: 16140064.


22. Parsons CL, Greenberger M, Gabal L, Bidair M, Barme G. The role of urinary potassium in the pathogenesis and diagnosis of interstitial cystitis. J Urol 1998;159:1862-1866. PMID: 9598476.


23. Ryan D, Robards K, Prenzler PD, Kendall M. Recent and potential developments in the analysis of urine: a review. Anal Chim Acta 2011;684:8-20. PMID: 21167980.


24. Keay SK, Szekely Z, Conrads TP, Veenstra TD, Barchi JJ Jr, Zhang CO, et al. An antiproliferative factor from interstitial cystitis patients is a frizzled 8 protein-related sialoglycopeptide. Proc Natl Acad Sci U S A 2004;101:11803-11808. PMID: 15282374.



25. Keay S, Zhang CO, Kagen DI, Hise MK, Jacobs SC, Hebel JR, et al. Concentrations of specific epithelial growth factors in the urine of interstitial cystitis patients and controls. J Urol 1997;158:1983-1988. PMID: 9334654.


26. Keay S, Zhang CO, Hise MK, Hebel JR, Jacobs SC, Gordon D, et al. A diagnostic in vitro urine assay for interstitial cystitis. Urology 1998;52:974-978. PMID: 9836539.


27. Keay S, Zhang CO, Marvel R, Chai T. Antiproliferative factor, heparin-binding epidermal growth factor-like growth factor, and epidermal growth factor: sensitive and specific urine markers for interstitial cystitis. Urology 2001;57(6 Suppl 1):104. PMID: 11378066.

28. Kim J, Keay SK, Dimitrakov JD, Freeman MR. p53 mediates interstitial cystitis antiproliferative factor (APF)-induced growth inhibition of human urothelial cells. FEBS Lett 2007;581:3795-3799. PMID: 17628545.



29. Keay S. Cell signaling in interstitial cystitis/painful bladder syndrome. Cell Signal 2008;20:2174-2179. PMID: 18602988.


30. Kim J, Keay SK, Freeman MR. Heparin-binding epidermal growth factor-like growth factor functionally antagonizes interstitial cystitis antiproliferative factor via mitogen-activated protein kinase pathway activation. BJU Int 2009;103:541-546. PMID: 18990151.



31. Kim J, Freeman MR. Antiproliferative factor signaling and interstitial cystitis/painful bladder syndrome. Int Neurourol J 2011;15:184-191. PMID: 22259731.



32. Yang W, Chung YG, Kim Y, Kim TK, Keay SK, Zhang CO, et al. Quantitative proteomics identifies a beta-catenin network as an element of the signaling response to Frizzled-8 protein-related antiproliferative factor. Mol Cell Proteomics 2011;10:M110.007492.



33. Kim J, Ji M, DiDonato JA, Rackley RR, Kuang M, Sadhukhan PC, et al. An hTERT-immortalized human urothelial cell line that responds to anti-proliferative factor. In Vitro Cell Dev Biol Anim 2011;47:2-9. PMID: 21136194.



34. Wishart DS. Advances in metabolite identification. Bioanalysis 2011;3:1769-1782. PMID: 21827274.


35. Serkova NJ, Brown MS. Quantitative analysis in magnetic resonance spectroscopy: from metabolic profiling to in vivo biomarkers. Bioanalysis 2012;4:321-341. PMID: 22303835.


36. Hughes C, Brown M, Clemens G, Henderson A, Monjardez G, Clarke NW, et al. Assessing the challenges of Fourier transform infrared spectroscopic analysis of blood serum. J Biophotonics 2014;7:180-188. PMID: 24488587.


37. Heemskerk AA, Deelder AM, Mayboroda OA. CE-ESI-MS for bottom-up proteomics: Advances in separation, interfacing and applications. Mass Spectrom Rev 2014 5 23;[Epub]. http://dx.doi.org/10.1002/mas.21432.

38. Robledo VR, Smyth WF. Review of the CE-MS platform as a powerful alternative to conventional couplings in bio-omics and target-based applications. Electrophoresis 2014;35:2292-2308. PMID: 24488766.


39. Zheng C, Zhang S, Ragg S, Raftery D, Vitek O. Identification and quantification of metabolites in (1)H NMR spectra by Bayesian model selection. Bioinformatics 2011;27:1637-1644. PMID: 21398670.



40. Krumsiek J, Suhre K, Evans AM, Mitchell MW, Mohney RP, Milburn MV, et al. Mining the unknown: a systems approach to metabolite identification combining genetic and metabolic information. PLoS Genet 2012;8:e1003005. PMID: 23093944.



41. Zhou B, Wang J, Ressom HW. MetaboSearch: tool for mass-based metabolite identification using multiple databases. PLoS One 2012;7:e40096. PMID: 22768229.



42. You S, Yang W, Anger JT, Freeman MR, Kim J. 'Omics' approaches to understanding interstitial cystitis/painful bladder syndrome/bladder pain syndrome. Int Neurourol J 2012;16:159-168. PMID: 23346481.



43. Yoshimura N. Editorial comment from Dr. Yoshimura to Potential urine and serum biomarkers for patients with bladder pain syndrome/interstitial cystitis. Int J Urol 2014;21(Suppl 1):42. PMID: 24807493.


44. Hanno P. Editorial comment from Dr. Hanno to Potential urine and serum biomarkers for patients with bladder pain syndrome/interstitial cystitis. Int J Urol 2014;21(Suppl 1):41. PMID: 24807492.


45. Kuo HC. Potential urine and serum biomarkers for patients with bladder pain syndrome/interstitial cystitis. Int J Urol 2014;21(Suppl 1):34-41. PMID: 24807491.


46. Zhang A, Sun H, Wang X. Serum metabolomics as a novel diagnostic approach for disease: a systematic review. Anal Bioanal Chem 2012;404:1239-1245. PMID: 22648167.


47. Trivedi DK, Iles RK. Do not just do it, do it right: urinary metabolomics -establishing clinically relevant baselines. Biomed Chromatogr 2014 5 02;[Epub]. http://dx.doi.org/10.1002/bmc.3219.

48. Zhang S, Nagana Gowda GA, Ye T, Raftery D. Advances in NMR-based biofluid analysis and metabolite profiling. Analyst 2010;135:1490-1498. PMID: 20379603.



49. Gebregiworgis T, Powers R. Application of NMR metabolomics to search for human disease biomarkers. Comb Chem High Throughput Screen 2012;15:595-610. PMID: 22480238.


50. Kohl SM, Klein MS, Hochrein J, Oefner PJ, Spang R, Gronwald W. State-of-the art data normalization methods improve NMR-based metabolomic analysis. Metabolomics 2012;8(Suppl 1):146-160. PMID: 22593726.



51. Veselkov KA, Vingara LK, Masson P, Robinette SL, Want E, Li JV, et al. Optimized preprocessing of ultra-performance liquid chromatography/mass spectrometry urinary metabolic profiles for improved information recovery. Anal Chem 2011;83:5864-5872. PMID: 21526840.


52. Van QN, Klose JR, Lucas DA, Prieto DA, Luke B, Collins J, et al. The use of urine proteomic and metabonomic patterns for the diagnosis of interstitial cystitis and bacterial cystitis. Dis Markers 2003-2004;19:169-183. PMID: 15258332.




53. Fukui Y, Kato M, Inoue Y, Matsubara A, Itoh K. A metabonomic approach identifies human urinary phenylacetylglutamine as a novel marker of interstitial cystitis. J Chromatogr B Analyt Technol Biomed Life Sci 2009;877:3806-3812.


54. Landis JR, Williams DA, Lucia MS, Clauw DJ, Naliboff BD, Robinson NA, et al. The MAPP research network: design, patient characterization and operations. BMC Urol 2014;14:58. PMID: 25085119.



55. Clemens JQ, Mullins C, Kusek JW, Kirkali Z, Mayer EA, Rodriguez LV, et al. The MAPP research network: a novel study of urologic chronic pelvic pain syndromes. BMC Urol 2014;14:58. PMID: 25085119.



Fig. 1
A workflow for interstitial cystitis (IC) metabolic profiling. NMR, nuclear magnetic resonance; GC-MS, gas chromatography-mass spectrometry; LC-MS, liquid chromatography-mass spectrometry; FT-IR, fourier transform infrared spectrometers; PCA, principal component analysis; PLS-DA, partial least squares discriminant analysis; OPLS-DA, orthogonal partial least squares discriminant analysis; ID, identification.